Yaml header rmarkdown4/1/2023  To fine control the formatting of citations and reference lists, users want to consult thisĪlso, for general reference management and obtaining references in Bibtex format PaperpileĬan be very helpful. Special section called References needs to be specified at the end of the R Markdown script. Reference to a reference list at the end of the output document. This will place the citation inline in the text and add the corresponding (2013), one uses its reference identifier ( e.g. Reference identifier present in the Bibtex database listed in the metadata section Script, one uses the syntax where needs to be replaced with a Reference collections should be stored in a separateįile in Bibtex or other supported formats. Citations and bibliographiesĬitations and bibliographies can be autogenerated in R Markdown in a similar This Wikibooks tutorial, or use an onlineĮquation rendering and checking tool, such as this one. To learn LaTeX syntax for mathematical equations, one can consult various online manuals, such as They have been added to print the code chunk syntax in this document. The following shows an example of such a code chunk. A code chunk is terminated by a new line starting with three backticks. R Code Chunks can be embedded in an R Markdown script by using three backticksĪt the beginning of a new line along with arguments enclosed in curly bracesĬontrolling the behavior of the code. To execute the Makefile, run the following To apply it to a custom Rmd file, one needs open the Makefile in a textĮditor and change the value assigned to MAIN (line 13) to the base name of A sample Makefile for rendering the above sample.Rmd can be $ Rscript -e "rmarkdown::render('sample.Rmd', output_format='BiocStyle::html_document', clean=TRUE)"Īlternatively, one can use a Makefile to evaluate and render an R Markdown To render to PDF format, use the argument setting: output_format="pdf_document".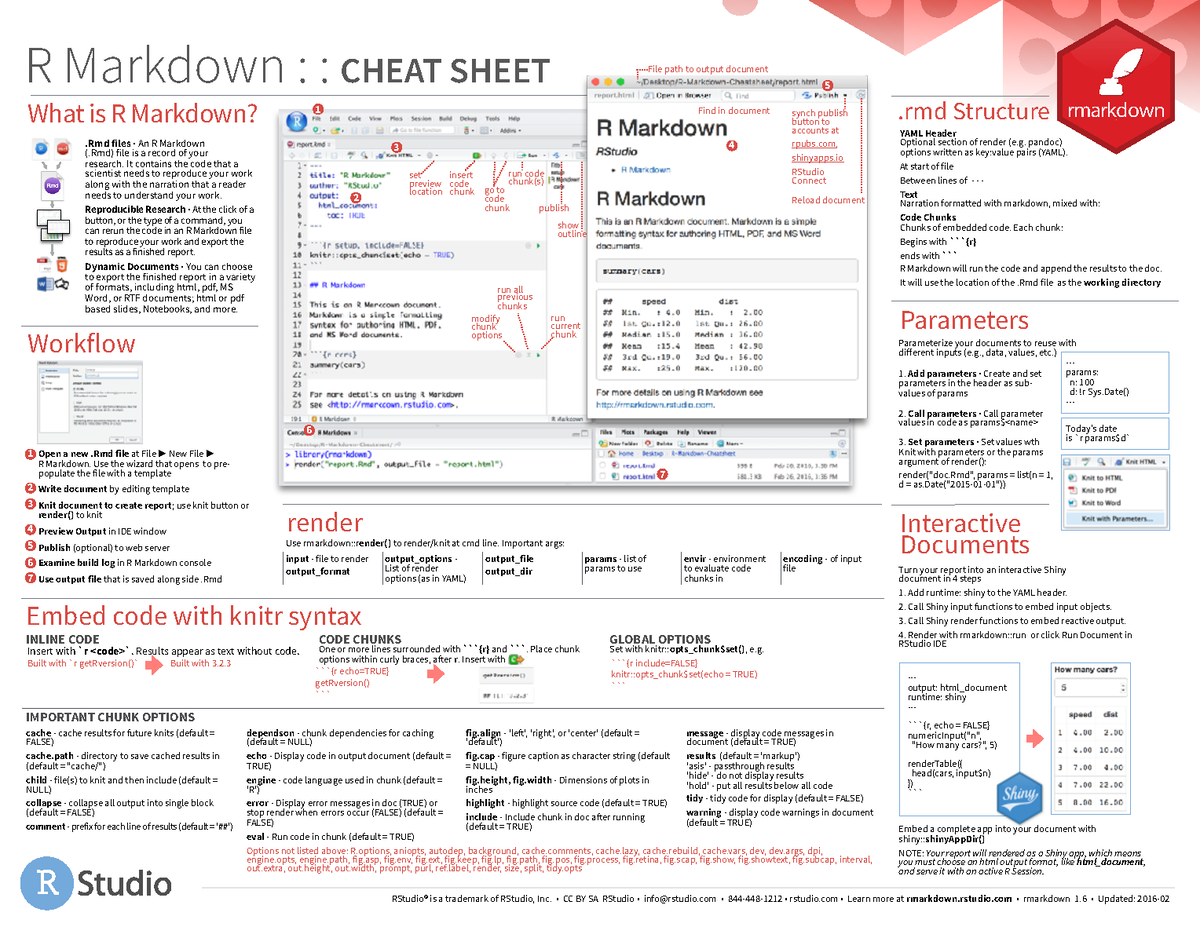  
The following shows two options how to run the rendering from the command-line. rmarkdown::render("sample.Rmd", clean=TRUE, output_format="BiocStyle::html_document") Alternatively, one can specify several output formats in the metadata section. The setting output_format="all" will generateĪll supported output formats. The output_format argument defines the format of the output ( e.g. The BiocStyle:: prefix will use theĪn R Markdown script can be evaluated and rendered with the following render command or by pressing the knit button in RStudio. With pdf_document and html_document, respectively. For instance, PDF and HTML output can be defined Options for customizing the output format. The metadata section also includes both title, author, and date information as well as The metadata section (YAML header) in an R Markdown script defines how it will be processed and RStudio, vim or emacs), initilize an R session and then direct their R session to Users want to download these files, open the sample.Rmd file with their preferred R IDE
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
